Dr. Endy and his team have developed a cellular logic gate, dubbed a transcriptor, which was the last of three things necessary to making cellular computing a reality: a way to store information, a method of transporting information, and something to perform basic logic operations. Storage has been achieved through recoding DNA into storing massive amounts of data. By re-configuring a harmless virus, data is able to be transferred between different cells and components in a cell. Lastly Endy’s transcriptor is able to process basic logic. Though in the infant stages of research, the possibilities of these breakthroughs include combating diseases and making livestock and crops safer.
Introduction
Thirty miles outside of Shanghai,China, a railway manufacturing plant spills a highly dangerous chemical that seeps into a river that local farmers depend on to irrigate their crops. In a couple days these crops turn an iridescent blue, alerting the farmers to the contamination. Halfway around the world, an absent minded man looks down at his phone at a notice that popped up, alerting him of low glucose levels, and averting what could have been a life threatening event.
What do these two seemingly disparate events have in common? They’re both scenarios that can be made possible with biological computing, utilizing an organism’s cells to perform many of the same tasks that modern day computers can. Transcriptors in the crops (the direct parallel being transistors in most modern day electronics) detect the chemical inside the plant and when it reaches a threshold limit, releases RNA Polymerase along strands of DNA, causing the gene that controls the color to express itself as blue rather than its standard color. Similarly for the example with the man, transcriptors monitor the flow of glucose molecules in the blood stream. This information can be stored in strands of DNA, whose sole purpose is the storage of data, and then accessed by a computing device that’s built to be integrated with these new synthetically engineered computing systems, allowing the collection of data so that corrections can be made upon the results. Theoretically these systems could even eliminate the need human correction; going off of the example of the man with low glucose levels, the cellular computer could then deliver specific amounts of insulin into the bloodstream.
Background
Electronic devices that perform computations, “computers”, consist of three main things: a way to store information , a way to transfer information between different parts, and the ability to process a basic system of logic [1]. Likewise if a biological system wants to act as a computational device it would have to have these same three things. Relatively early on it has been demonstrated that it is easy to store information by encoding DNA in a certain way [2], and the transferring of information has been done by re-configuring certain non-harmful viruses [3]. The missing link was a transistor like operator that could process basic logic, which has eluded researchers for a while.
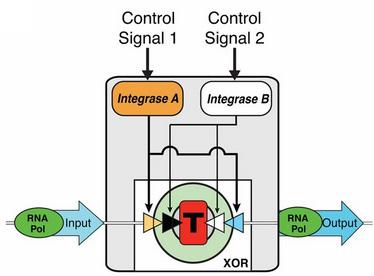
Recently, biological engineers led by Drew Endy at Stanford University have made that final breakthrough, developing a “transistor” (that they have named transcriptor) made of DNA and RNA that is capable of acting as logic gates, such as AND and XOR (shown in Fig. 1) [4]. Now with all three components in place, further proof of concept and initial research should start to emerge, paving the way for many future possibilities.
Recently, biological engineers led by Drew Endy at Stanford University have made that final breakthrough, developing a “transistor” (that they have named transcriptor) made of DNA and RNA that is capable of acting as logic gates, such as AND and XOR (shown in Fig. 1) [4]. Now with all three components in place, further proof of concept and initial research should start to emerge, paving the way for many future possibilities.
Breaking Down the Components
Storage
If you’re currently reading this online, then the computer or internet enabled device that you have before you possess some form of “memory” (Hard drive, RAM). The tasks that are performed are stored and are able to be accessed during a later time, anything from storing your favorite photos to looking at an essay you wrote a year ago.
When researchers started to look at a way to replicate this in living systems, they ran into a fortunate coincidence, such a storage method already existed, utilizing deoxyribonucleic acid (DNA), ribonucleic acid (RNA), and various other proteins. These elements are combined into sequences that are later read and used by ribosomes to produce a certain type of protein that the cell needs. Scientists have figured out ways to produce their own strands of DNA encoded in a way that translates into something like an image or a word.
Using DNA as a storage method not only works, but presents many benefits as well. DNA storage is very dense, supporting EXAbytes (〖10〗^6 terabytes!) per gram of single stranded DNA , is already used by life which allows for easy exchange between the cellular computers and the organism, and lasts over a millennia even in non-ideal states [5].
Using a very specific type of printer, DNA designers can produce DNA strands in a strand that represents a sequence of information [6]. This can be then “uploaded” to the cell that contains our cellular computer. Or if the device itself is producing the information, then it can write the data to strands of DNA directly [7] that later we can access.
When researchers started to look at a way to replicate this in living systems, they ran into a fortunate coincidence, such a storage method already existed, utilizing deoxyribonucleic acid (DNA), ribonucleic acid (RNA), and various other proteins. These elements are combined into sequences that are later read and used by ribosomes to produce a certain type of protein that the cell needs. Scientists have figured out ways to produce their own strands of DNA encoded in a way that translates into something like an image or a word.
Using DNA as a storage method not only works, but presents many benefits as well. DNA storage is very dense, supporting EXAbytes (〖10〗^6 terabytes!) per gram of single stranded DNA , is already used by life which allows for easy exchange between the cellular computers and the organism, and lasts over a millennia even in non-ideal states [5].
Using a very specific type of printer, DNA designers can produce DNA strands in a strand that represents a sequence of information [6]. This can be then “uploaded” to the cell that contains our cellular computer. Or if the device itself is producing the information, then it can write the data to strands of DNA directly [7] that later we can access.
Transmission
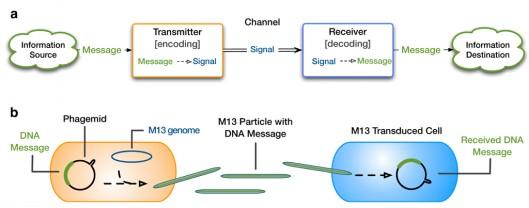
Though a description for a method of storage was described, if that information is static and wasn’t able to be accessed by both humans and other parts of the cellular computer, it would be useless and gather dust, so to speak. In a traditional computer, this transfer is accomplished by buses. In our cellular computer this was trickier than the problem of storage, as the DNA solution was obvious. Finally after digging though researchers at Stanford University were able to reprogram a virus, M13, to perform this task [3]. Normally the virus acts as a parasite by feeding off cellular “food” and encapsulating loose strands of DNA in protein to send to other cells to infect them with this repackaged DNA. Drew Endy and Monica Ortiz have been able to utilize their DNA repackaging abilities to transfer messages of their own choosing to a specified destination, in essence creating the possibility for a giant network of information flow to exist between individual “computer” parts and separate synthetic biological devices located in a comparatively far off location [3] . Fig. 2 shows a traditional electronic computational device’s method of information transfer and the analog of the system in living creature.
Bonnet/Endy/ Stanford University School of Engineering
Figure 2: Transfer of information in a computer and the biological
equivalent of the process.
equivalent of the process.
Processing a basic system of logic
The final key that was eluding scientists that would complete the cellular computer was a way to process a basic system of logic. In common english, a basic system of logic tests to see if a certain set of conditions is true or not and then react based on the results. If you use the logic operator AND for example, the output is set as True only if condition A and condition B are met via the inputs. The output may be used as a counting method, telling you how many elements meet your test, and recording the number of True or False, or the output might be a direct action. Imagine a common door acts as a logic gate that employs the AND operator and we set the conditions to having a key and turning a knob. If both of these conditions are met, then door represents this True value by opening and allowing you inside. If the conditions are not met, then you’ve generated a False value and the door will remain closed.
Dr. Endy and a team of researchers have found a way to employ these logic operators with cellular modules built from plasmids and integrases [4]. These modules contain Boolean integrase logic gates( BIL gates), that are capable of applying tests of logic. These gates can be further configured through control signals that switch the parameters of their test. This breakthrough acts in way similar to transistors, which is why these particular modules are termed “transcriptors”, a combination of transistor and scripting, as in re-scripting DNA sequences by releasing enzymes depending on the output value. You can see an example of this in Fig. 1.
Dr. Endy and a team of researchers have found a way to employ these logic operators with cellular modules built from plasmids and integrases [4]. These modules contain Boolean integrase logic gates( BIL gates), that are capable of applying tests of logic. These gates can be further configured through control signals that switch the parameters of their test. This breakthrough acts in way similar to transistors, which is why these particular modules are termed “transcriptors”, a combination of transistor and scripting, as in re-scripting DNA sequences by releasing enzymes depending on the output value. You can see an example of this in Fig. 1.
Future Potential
Of course, the field of synthetic biological computing is just beginning to grow and the three basic parts necessary to computation (Storage, Transfer, and Logic Processing) that have been demonstrated as implementable in organisms serve more as proof of concepts than perfected technology. With that being said though, the potential for these discoveries are immense. The examples at the beginning of the paper are just the tip of the iceberg.
At the very simplest end of the spectrum of opportunities that this technology affords is simple counting tasks. This alone is stunning. Having current information from your body, or an animals’ or plants’, allows doctors/ veterinarians/ botanists to make educated, informed decisions based on instant data, which allows them to catch or prevent many diseases before they start. Using a transcriptor utilizing an AND operator could register True values if two certain proteins are present that are associated with a particular disease [9]. A doctor can be alerted by these computers that you’re starting to have abnormal levels of these two proteins and he can make steps to remedy the situation. Of course, you can eliminate the doctor from the equation altogether by allowing the information gathered from the AND transcriptor to be transmitted to another transcriptor via the M13 virus, and that transcriptor can tell a cell to produce chemicals to counteract the rise in harmful protein levels.
Or you might install these computers in a plant to not only react to harmful chemicals by turning blue, but to a variety of environmental conditions and respond by altering the plant to be able to thrive in that environment, for example, being able to repel insects or producing an extra layer of foliage to insulate itself from extreme cold.
At the very simplest end of the spectrum of opportunities that this technology affords is simple counting tasks. This alone is stunning. Having current information from your body, or an animals’ or plants’, allows doctors/ veterinarians/ botanists to make educated, informed decisions based on instant data, which allows them to catch or prevent many diseases before they start. Using a transcriptor utilizing an AND operator could register True values if two certain proteins are present that are associated with a particular disease [9]. A doctor can be alerted by these computers that you’re starting to have abnormal levels of these two proteins and he can make steps to remedy the situation. Of course, you can eliminate the doctor from the equation altogether by allowing the information gathered from the AND transcriptor to be transmitted to another transcriptor via the M13 virus, and that transcriptor can tell a cell to produce chemicals to counteract the rise in harmful protein levels.
Or you might install these computers in a plant to not only react to harmful chemicals by turning blue, but to a variety of environmental conditions and respond by altering the plant to be able to thrive in that environment, for example, being able to repel insects or producing an extra layer of foliage to insulate itself from extreme cold.
Conclusion
Advances in logic based cellular modules and combining these advances with existing methods of information storage and transfer makes the concept of cellular computing a reality, with many future possibilities. They are not meant to replace traditional computers, but to be used in conjunction of them to give us more information and control in living systems. Possible uses include monitoring the human body for the prevention of disease and harm and improving the hardiness of livestock and crops in a medley of situations [8].
References
-
- [1] Sebastein Anthony (March 29, 2013). “Stanford creates biological transistors, the final step towards computers inside living cells”. Extreme Tech.
- [2] Gautam Naik. (2013, Jan 24). Storing Digital Data in DNA [Online]. Available: http://online.wsj.com/article/SB10001424127887324539304578259883507543150.html
- [3] Monica E Ortiz, Drew Endy, “Engineered cell-cell communication via DNA
- messaging,” Journal of Biological Engineering 2012, 6:16
- [4] Jerome Bonnet, Peter Yin*, Monica E. Ortiz, Pakpoom Subsoontorn, Drew Endy, “Amplifying Genetic Logic Gates,” Department of Bioengineering. Stanford Univ., Stanford,Ca. Rep. 10.1126/science.1232758, 2013
- [5] George M. Church, Yuan Gao, Sriram Kosuri , “Next-Generation Digital Information
- Storage in DNA,” U.S. Office of Naval Research 10.1126/science.1226355, 2012
- [6] Emily M. LeProust, Bill J. Peck,Konstantin Spirin, Heather Brummel McCuen, “Synthesis of high-quality libraries of long (150mer) oligonucleotides by a novel depurination controlled process,” 2010 March 20. doi: 10.1093/nar/gkq163
- [7] Devin Burill, Pamela Silver ,“Making Cellular Memories” 10.1016/j.cell.2009.12.034 Cell, Volume 140, Issue 1, 13-18, 8 January 2010
- [8] Yaakov Benenson, “Biomolecular computing systems: principles, progress and potential”, (July 2012) doi:10.1038/nrg3197